 |
|
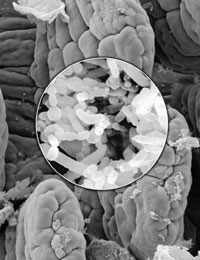
Scanning electron microscope images of B. thetaiotaomicron, a prominent human gut bacterium, and the intestine.
|
A human being can be viewed as a biological eco-system that includes distinctly human cells and tissues as well as thousands of microbial species that co-exist in various parts of the body.
In fact, "within the body of a healthy adult, microbial cells are estimated to outnumber human cells by a factor of ten to one. These communities, however, remain largely unstudied, leaving almost entirely unknown their influence upon human development, physiology, immunity, and nutrition. To take advantage of recent technological advances and to develop new ones, the NIH Roadmap has initiated the Human Microbiome Project (HMP) with the mission of generating resources enabling comprehensive characterization of the human microbiota and analysis of its role in human health and disease." [https://commonfund.nih.gov/hmp]
The BCM-HGSC, with Genome Sequencing Center at Washington University, the Broad Institute of MIT and Harvard, and the J. Craig Venter Institute addressed some early goals of this project. The HMP metagenomic sample sequencing compared microbial communities between individuals, body sites, and states (e.g., disease, diet, age).1 Pilot projects determined the appropriate sequencing platforms, quality controls, and annotation pipelines for generating reference genomes3 and sequencing metagenomic samples.2
Resources
| Entry # | Reference strains | Center |
|---|---|---|
| 1 | Acidaminococcus sp. 2_2_8 | Broad |
| 2 | Acidovorax AM084006 | JCVI |
| 3 | Acinetobacter haemolyticus | JCVI |
| 4 | Acinetobacter schindleri | BCM-HGSC |
| 5 | Acinetobacter sp. ATCC#27244 | BCM-HGSC |
| 6 | Actinobacterium M28 | BCM-HGSC |
| 7 | Actinomyces coleocanis | BCM-HGSC |
| 8 | Actinomyces naeslundii | JCVI |
| 9 | Actinomyces urogenitalis | BCM-HGSC |
| 10 | Aerococcus christensenii | BCM-HGSC |
| 11 | Aeromicrobium marinum | BCM-HGSC |
| 12 | Alistipes putredinis DSM 17216 | WUGSC |
| 13 | Alistipes sp. 1_2_47A | Broad |
| 14 | Alistipes sp. 10_1_37_FAA | Broad |
| 15 | Allochromatium vinosum | BCM-HGSC |
| 16 | Anaerococcus hydrogenalis DSM 7454 | WUGSC |
| 17 | Anaerococcus lactolyticus | BCM-HGSC |
| 18 | Anaerococcus lactolyticus (additional) | BCM-HGSC |
| 19 | Anaerococcus lactolyticus ATCC#51172 | BCM-HGSC |
| 20 | Anaerococcus tetradius | JCVI |
| 21 | Anaerococcus tetradius | BCM-HGSC |
| 22 | Anaerofustis stercorihominis DSM 17244 | WUGSC |
| 23 | Anaerostipes caccae DSM14662 | WUGSC |
| 24 | Anaerotruncus colihominis DSM 17241 | WUGSC |
| 25 | Aneurinibacillus aneurinolyticus DSM 5562 | WUGSC |
| 26 | Arcobacter cryaerophilus ATCC 49615 | JCVI |
| 27 | Atopobium vaginae | JCVI |
| 28 | Atopobium vaginae ATCC#BAA-55 | BCM-HGSC |
| 29 | Bacteroides pectinophilus ATCC 43243 | WUGSC |
| 30 | Bacteroides coprocola DSM 17136 | WUGSC |
| 31 | Bacteroides dorei DSM 17855 | WUGSC |
| 32 | Bacteroides eggerthii DSM 20697 | WUGSC |
| 33 | Bacteroides finegoldii DSM 17565 | WUGSC |
| 34 | Bacteroides galacturonicus ATCC 43244 | WUGSC |
| 35 | Bacteroides intestinalis DSM 17393 | WUGSC |
| 36 | Bacteroides plebeius DSM 17135 | WUGSC |
| 37 | Bacteroides heparinolytica | JCVI |
| 38 | Bacteroides merdae ATCC 43184 | WUGSC |
| 39 | Bacteroides sp. 1_1_11 | Broad |
| 40 | Bacteroides sp. 1_1_14 | Broad |
| 41 | Bacteroides sp. 1_1_30 | Broad |
| 42 | Bacteroides sp. 1_1_6 | Broad |
| 43 | Bacteroides sp. 2_1_16 | Broad |
| 44 | Bacteroides sp. 2_1_22 | Broad |
| 45 | Bacteroides sp. 2_1_33B | Broad |
| 46 | Bacteroides sp. 2_1_7 | Broad |
| 47 | Bacteroides sp. 2_2_4 | Broad |
| 48 | Bacteroides sp. 20_3 | Broad |
| 49 | Bacteroides sp. 3_1_12 | Broad |
| 50 | Bacteroides sp. 3_1_13 | Broad |
| 51 | Bacteroides sp. 3_1_19 | Broad |
| 52 | Bacteroides sp. 3_1_23 | Broad |
| 53 | Bacteroides sp. 3_1_33FAA | Broad |
| 54 | Bacteroides sp. 3_1_40A | Broad |
| 55 | Bacteroides sp. 3_2_5 | Broad |
| 56 | Bacteroides sp. 4_1_36 | Broad |
| 57 | Bacteroides sp. 4_3_47FAA | Broad |
| 58 | Bacteroides sp. 9_1_42FAA | Broad |
| 59 | Bacteroides stercoris ATCC 43183 | WUGSC |
| 60 | Bacteroides uniformis DSM 6597 | WUGSC |
| 61 | Bdellovibrio AF148938 | JCVI |
| 62 | Bifidobacterium dentium ATCC 27678 | WUGSC |
| 63 | Bifidobacterium angulatum DSM 20098 | WUGSC |
| 64 | Bifidobacterium bifidum DSM 20456 | WUGSC |
| 65 | Bifidobacterium breve DSM 20213 | WUGSC |
| 66 | Bifidobacterium catenulatum DSM 16992 | WUGSC |
| 67 | Bifidobacterium gallicum DSM 20093 | WUGSC |
| 68 | Bifidobacterium bifidum NCIMB 41171 | Broad |
| 69 | Bifidobacterium dentium | JCVI |
| 70 | Bifidobacterium longum infantis strain 52484 | Broad |
| 71 | Bifidobacterium pseudocatenulatum DSM 20438 | WUGSC |
| 72 | Bifidobacterium sp. 12_1_47BFAA | Broad |
| 73 | Brachybacterium sp.C3H-211 | JCVI |
| 74 | Brevibacterium mcbrellneri ATCC#49030 | BCM-HGSC |
| 75 | Brevibacterium paucivorans | JCVI |
| 76 | Brevundimonas aurantiaca | JCVI |
| 77 | Bryantella formatexigens DSM 14469 | WUGSC |
| 78 | Butyrivibrio crossotus DSM 2876 | WUGSC |
| 79 | Campylobacter Bacterioides ureolyticus ATCC 33387 (type) | JCVI |
| 80 | Capnocytophaga gingivalis | JCVI |
| 81 | Carnobacterium AJ427446 | JCVI |
| 82 | Catenibacterium mitsuokai DSM 15897 | WUGSC |
| 83 | Cedecea davisae DSM 4568 | WUGSC |
| 84 | Cetobacterium somerae ATCC N/A | WUGSC |
| 85 | Chitinophaga AJ318177 | JCVI |
| 86 | Chryseobacterium gleum ATCC#35910 | BCM-HGSC |
| 87 | Citrobacter youngae ATCC29220 | WUGSC |
| 88 | Citrobacter sp. 30_2 | Broad |
| 89 | Clostridium celatum DSM 1785 | WUGSC |
| 90 | Clostridium sporogenes ATCC 15579 | WUGSC |
| 91 | Clostridium asparagiforme DSM 15981 | WUGSC |
| 92 | Clostridium bartlettii DSM 16795 | WUGSC |
| 93 | Clostridium celatum DSM 1785 | WUGSC |
| 94 | Clostridium hathewayi DSM 13479 | WUGSC |
| 95 | Clostridium hiranonis DSM 13275 | WUGSC |
| 96 | Clostridium hylemonae DSM 15053 | WUGSC |
| 97 | Clostridium leptum DSM 753 | WUGSC |
| 98 | Clostridium methylpentosum DSM 5476 | WUGSC |
| 99 | Clostridium nexile DSM 1787 | WUGSC |
| 100 | Clostridium nexile-related A2-232 | WUGSC |
| 101 | Clostridium orbiscindens DSM 6740 | WUGSC |
| 102 | Clostridium scatologenes DSM 757 | WUGSC |
| 103 | Clostridium scindens ATCC 35704 | WUGSC |
| 104 | Clostridium sp. 1_1_41A1FAA | Broad |
| 105 | Clostridium boltaea ATCC BAA-613 | WUGSC |
| 106 | Clostridium eutactus-related L2-50 | WUGSC |
| 107 | Clostridium indolis-related SS2/1 | WUGSC |
| 108 | Clostridium inocuum 6_1_30 | Broad |
| 109 | Clostridium ramosum DSM 1402 | WUGSC |
| 110 | Clostridium saccharolyticum-related M62/1 | WUGSC |
| 111 | Clostridium sp A2-166 | WUGSC |
| 112 | Clostridium sp DSM 4029 | WUGSC |
| 113 | Clostridium sp SR1/1 | WUGSC |
| 114 | Clostridium sp. ATCC BAA-442 | WUGSC |
| 115 | Clostridium sp. 7_2_43FAA | Broad |
| 116 | Clostridium spiroforme DSM 1552 | WUGSC |
| 117 | Clostridium symbiosum ATCC 14940 | WUGSC |
| 118 | Collinsella intestinalis DSM 13280 | WUGSC |
| 119 | Collinsella stercoris DSM 13279 | WUGSC |
| 120 | Coprobacillus sp. 29_1 | Broad |
| 121 | Coprobacillus sp. 8_1_38FAA | Broad |
| 122 | Coprobacillus sp. 8_2_54BFAA | Broad |
| 123 | Coprococcus comes ATCC 27758 | WUGSC |
| 124 | Coprococcus eutactus ATCC 27759 | WUGSC |
| 125 | Corynebacterium ammoniagenes DSM 20306 | WUGSC |
| 126 | Corynebacterium accolens | JCVI |
| 127 | Corynebacterium accolens ATCC#49725 | BCM-HGSC |
| 128 | Corynebacterium aurimucosum | BCM-HGSC |
| 129 | Corynebacterium efficiens | BCM-HGSC |
| 130 | Corynebacterium genitalium ATCC#33030 | BCM-HGSC |
| 131 | Corynebacterium glutamicum | BCM-HGSC |
| 132 | Corynebacterium jelkelum | BCM-HGSC |
| 133 | Corynebacterium lipophiloflavum | BCM-HGSC |
| 134 | Corynebacterium nigricans ATCC#700975 | BCM-HGSC |
| 135 | Corynebacterium pseudogenitalium ATCC#33035 | BCM-HGSC |
| 136 | Corynebacterium seminale ATCC#51866 | BCM-HGSC |
| 137 | Corynebacterium seminale ATCC#51867 | BCM-HGSC |
| 138 | Corynebacterium striatum | BCM-HGSC |
| 139 | Corynebacterium sundsvallense | BCM-HGSC |
| 140 | Cryptobacterium curtum1 | JCVI |
| 141 | Deinococcus AE002076 | JCVI |
| 142 | Delftia acidovorans1 | JCVI |
| 143 | Desulfomicrobium orale | JCVI |
| 144 | Desulfovibrio fairfieldensis4 | JCVI |
| 145 | Desulfovibrio piger DSM 749 | WUGSC |
| 146 | Desulfovibrio sp. 3_1_syn3 | Broad |
| 147 | Dialister invisus | JCVI |
| 148 | Diaphorobacter nitroreducens | JCVI |
| 149 | Dietzia sp. E9_21 | JCVI |
| 150 | Dorea formicigenerans ATCC 27755 | WUGSC |
| 151 | Dorea sp. 1_1_29AFAA | Broad |
| 152 | Dorea sp. 4_1_36FAA | Broad |
| 153 | Edwardsiella tarda ATCC 23685 | WUGSC |
| 154 | Enterobacter cancerogenus ATCC 35316 | WUGSC |
| 155 | Enterobacter asburiae | JCVI |
| 156 | Enterococcus faecalis | JCVI |
| 157 | Enterococcus faecalis ATCC#29200 | BCM-HGSC |
| 158 | Enterococcus faecalis HH22 | BCM-HGSC |
| 159 | Enterococcus faecalis OG1RF | BCM-HGSC |
| 160 | Enterococcus faecalis TX0104 | BCM-HGSC |
| 161 | Enterococcus faecalis TX1322 | BCM-HGSC |
| 162 | Enterococcus faecium DO | BCM-HGSC |
| 163 | Enterococcus faecium TX1330 | BCM-HGSC |
| 164 | Erysipelothrix rhusiopathiae | BCM-HGSC |
| 165 | Escherichia sp. 1_1_43 | Broad |
| 166 | Escherichia sp. 3_2_53FAA | Broad |
| 167 | Escherichia sp. 4_1_40B | Broad |
| 168 | Eschericia coli MG1655 | BCM-HGSC |
| 169 | Eschericia coli Trautner | BCM-HGSC |
| 170 | Eubacterium biforme DSM 3989 | WUGSC |
| 171 | Eubacterium cylindroides DSM 3983 | WUGSC |
| 172 | Eubacterium dolichum DSM 3991 | WUGSC |
| 173 | Eubacterium hadrum DSM 3319 | WUGSC |
| 174 | Eubacterium halii-related DSM 17630 | WUGSC |
| 175 | Eubacterium ramulus DSM 3995 | WUGSC |
| 176 | Eubacterium brachy4 | JCVI |
| 177 | Eubacterium hallii DSM 3353 | WUGSC |
| 178 | Eubacterium plautii ATCC 29863 | WUGSC |
| 179 | Eubacterium siraeum DSM 15702 | WUGSC |
| 180 | Eubacterium sp. 3_1_31 | Broad |
| 181 | Eubacterium sp. L2-7 | WUGSC |
| 182 | Faecalibacterium prausnitzii M21/2 | WUGSC |
| 183 | Faecalibacterium prausnitzii A2-165 | WUGSC |
| 184 | Falcivibrio vaginalis ATCC#43063 | BCM-HGSC |
| 185 | Finegoldia magna ATCC#53516 | BCM-HGSC |
| 186 | Fusobacterium prausnitzii DSM 17677 | WUGSC |
| 187 | Fusobacterium prausnitzii L2-6 | WUGSC |
| 188 | Fusobacterium gonidiaformans ATCC 25563 | Broad |
| 189 | Fusobacterium mortiferum ATCC 9817 | Broad |
| 190 | Fusobacterium nucleatum ATCC#10953 | BCM-HGSC |
| 191 | Fusobacterium nucleatum ATCC#23726 | BCM-HGSC |
| 192 | Fusobacterium sp 1_1_41FAA | Broad |
| 193 | Fusobacterium sp 11_3_2 | Broad |
| 194 | Fusobacterium sp. 12_1B | Broad |
| 195 | Fusobacterium sp. 2_1_31 | Broad |
| 196 | Fusobacterium sp. 2_1_37FAA | Broad |
| 197 | Fusobacterium sp. 21_1A | Broad |
| 198 | Fusobacterium sp. 3_1_27 | Broad |
| 199 | Fusobacterium sp. 3_1_33 | Broad |
| 200 | Fusobacterium sp. 3_1_36A2 | Broad |
| 201 | Fusobacterium sp. 3_1_5R | Broad |
| 202 | Fusobacterium sp. 4_1_13 | Broad |
| 203 | Fusobacterium sp. 4_8 | Broad |
| 204 | Fusobacterium sp. 7_1 | Broad |
| 205 | Fusobacterium ulcerans ATCC 49185, NCTC 12111 | Broad |
| 206 | Fusobacterium varium ATCC 27725 | Broad |
| 207 | Gardnerella vaginalis ATCC#14019 | BCM-HGSC |
| 208 | Gordonia sputi | JCVI |
| 209 | Grimontia hollisae DSM 15132 | WUGSC |
| 210 | Haemophilus AF224309 | JCVI |
| 211 | Haemophilus ducreyi ATCC#700724 | BCM-HGSC |
| 212 | Helicobacter canadensis | BCM-HGSC |
| 213 | Helicobacter canadensis MIT 98-5491 (LCDC 16143, ATCC 700968) | Broad |
| 214 | Helicobacter cinaedi ATCC BAA-847 | JCVI |
| 215 | Helicobacter cinaedi CCUG 18818 (NCTC 12423) | Broad |
| 216 | Helicobacter pullorum | BCM-HGSC |
| 217 | Helicobacter pullorum MIT 98-5489 (LCDC 15115) | Broad |
| 218 | Helicobacter pylori 35A | BCM-HGSC |
| 219 | Helicobacter pylori 74B | BCM-HGSC |
| 220 | Helicobacter rappini taxon 8 (ATCC 49309) | Broad |
| 221 | Helicobacter winghamensis (ATCC BAA-430) | Broad |
| 222 | Holdemania filiformis DSM 12042 | WUGSC |
| 223 | Hyphomicrobium facile | JCVI |
| 224 | Janibacter melonis | JCVI |
| 225 | Janthinobacterium lividum | JCVI |
| 226 | Klebsiella sp. 1_1_55 | Broad |
| 227 | Kocuria marina | JCVI |
| 228 | Lactobacillus brevis DSM 20054 | WUGSC |
| 229 | Lactobacillus acidophilus | BCM-HGSC |
| 230 | Lactobacillus amylolyticus | BCM-HGSC |
| 231 | Lactobacillus antri | BCM-HGSC |
| 232 | Lactobacillus coleohominis | BCM-HGSC |
| 233 | Lactobacillus crispatus | JCVI |
| 234 | Lactobacillus crispatus JVV01 | BCM-HGSC |
| 235 | Lactobacillus delbrueckii | BCM-HGSC |
| 236 | Lactobacillus gasseri JVV03 | BCM-HGSC |
| 237 | Lactobacillus helveticus | BCM-HGSC |
| 238 | Lactobacillus iners | BCM-HGSC |
| 239 | Lactobacillus jensenii JVV16 | BCM-HGSC |
| 240 | Lactobacillus johnsonii | BCM-HGSC |
| 241 | Lactobacillus lactis subsp. cremoris | BCM-HGSC |
| 242 | Lactobacillus paracasei 362.501388888889 | Broad |
| 243 | Lactobacillus reuteri CF48-3A | BCM-HGSC |
| 244 | Lactobacillus rhamnosus | BCM-HGSC |
| 245 | Lactobacillus sakei | BCM-HGSC |
| 246 | Lactobacillus suntoryeus | BCM-HGSC |
| 247 | Lactobacillus ultunensis | BCM-HGSC |
| 248 | Leptothrix sp. strain DhA-711 | JCVI |
| 249 | Leptotrichia amnionii | JCVI |
| 250 | Leuconostoc argentinum | JCVI |
| 251 | Leuconostoc mesenteroides | BCM-HGSC |
| 252 | Listeria grayi | BCM-HGSC |
| 253 | Megasphaera micronucliformis1 | JCVI |
| 254 | Mesorhizobium AP0030011 | JCVI |
| 255 | Methanobrevibacter smithii DSM 11975 | WUGSC |
| 256 | Methanobrevibacter oralis1 | JCVI |
| 257 | Methanobrevibacter smithii DSM 2374 | WUGSC |
| 258 | Methanobrevibacter smithii DSM 2375 | WUGSC |
| 259 | Methylobacillus AF289159 | JCVI |
| 260 | Methylobacterium AY741717 | JCVI |
| 261 | Micrococcus luteus4 | JCVI |
| 262 | Micromonas micros1 | JCVI |
| 263 | Mitsuokella multacida DSM 20544 | WUGSC |
| 264 | Mobiluncus curtisii | BCM-HGSC |
| 265 | Mobiluncus mulieris | BCM-HGSC |
| 266 | Mogibacterium timidum1 | JCVI |
| 267 | Moraxella osloensis4 | JCVI |
| 268 | Mycobacterium chlorophenolicum | JCVI |
| 269 | Mycobacterium parascrofulaceum ATCC#BAA-614 | BCM-HGSC |
| 270 | Mycoplasma fermentans ATCC#15474 | BCM-HGSC |
| 271 | Mycoplasma hominis ATCC#14268 | BCM-HGSC |
| 272 | Mycoplasma hominis ATCC#43519 | BCM-HGSC |
| 273 | Mycoplasma salivarum4 | JCVI |
| 274 | Mycoplasma spermatophilum ATCC#49695 | BCM-HGSC |
| 275 | Neisseria DQ409137 | JCVI |
| 276 | Neisseria gonorrhoeae ATCC#43070 | BCM-HGSC |
| 277 | Paenibacillus sp. | JCVI |
| 278 | Papillibacter sp. 4_1_38B | Broad |
| 279 | Pediococcus sp. 7_4A | Broad |
| 280 | Pedomicrobium australicum | JCVI |
| 281 | Peptostreptococcus anaerobius | JCVI |
| 282 | Peptostreptococcus micros ATCC 33270 | WUGSC |
| 283 | Phascolarctobacterium sp. 3_1_syn4 | Broad |
| 284 | Porphyromonas L16493 | JCVI |
| 285 | Prevotella AY207061 | JCVI |
| 286 | Prevotella bergensis | BCM-HGSC |
| 287 | Propionibacterium acnes | JCVI |
| 288 | Propionibacterium sp. 5_U_42AFAA | Broad |
| 289 | Proteus penneri ATCC 35198 | WUGSC |
| 290 | Proteus mirabilis | BCM-HGSC |
| 291 | Providencia rustigianii DSM 4541 | WUGSC |
| 292 | Providencia stuartii ATCC 25827 | WUGSC |
| 293 | Pseudomonas aeruginosa | JCVI |
| 294 | Rhodococcus corynebacterioides | JCVI |
| 295 | Rhodococcus equi | BCM-HGSC |
| 296 | Roseburia faecis DSM 16840 | WUGSC |
| 297 | Roseburia intestinalis DSM 14610 | WUGSC |
| 298 | Roseburia inulinivorans DSM 16841 | WUGSC |
| 299 | Roseburia inulinivorans L1-83 | WUGSC |
| 300 | Roseomonas cervicalis ATCC#49957 | BCM-HGSC |
| 301 | Ruminococcus callidus ATCC 27760 | WUGSC |
| 302 | Ruminococcus gnavus-related GM2/1 | WUGSC |
| 303 | Ruminococcus hansenii DSM 20583 | WUGSC |
| 304 | Ruminococcus luti DSM 14534 | WUGSC |
| 305 | Ruminococcus hydrogenotrophicus DSM 10507 | WUGSC |
| 306 | Ruminococcus lactaris ATCC 29176 | WUGSC |
| 307 | Ruminococcus sp. | JCVI |
| 308 | Ruminococcus sp. 4_1_47 FAA | Broad |
| 309 | Ruminococcus sp. 5_1_39AFAA | Broad |
| 310 | Ruminococcus sp. 5_1_39B FAA | Broad |
| 311 | Ruminococcus sp. 8_1_37 FAA | Broad |
| 312 | Selenomonas dianae2 | JCVI |
| 313 | Serratia liquefaciens | JCVI |
| 314 | Slackia exigua | JCVI |
| 315 | Sphingobacterium spiritivorum ATCC#33300 | BCM-HGSC |
| 316 | Sphingobacterium spiritivorum ATCC#33861 | BCM-HGSC |
| 317 | Sphingobium amiens | JCVI |
| 318 | Sphingopyxis AY328823 | JCVI |
| 319 | Staphylococcus aureus MN8 | BCM-HGSC |
| 320 | Staphylococcus aureus MRSA | BCM-HGSC |
| 321 | Staphylococcus aureus USA300 | BCM-HGSC |
| 322 | Staphylococcus capitis | JCVI |
| 323 | Staphylococcus epidermidis | BCM-HGSC |
| 324 | Stenotrophomonas maltophilia | JCVI |
| 325 | Streptococcus infantarius ATCC BAA-102 | WUGSC |
| 326 | Streptococcus agalactiae | JCVI |
| 327 | Streptococcus equinus | BCM-HGSC |
| 328 | Streptococcus pneumoniae | BCM-HGSC |
| 329 | Streptococcus sp. 2_1_36FAA | Broad |
| 330 | Subdoligranulum variabile DSM 15176 | WUGSC |
| 331 | Synergistes phyla sp. E1 E3_33 | JCVI |
| 332 | Synergistes sp. 3_1_syn1 | Broad |
| 333 | Thermomicrobium AY250886 | JCVI |
| 334 | Treponema amylovorum | JCVI |
| 335 | Tsukamurella tyrosinosolvens | JCVI |
| 336 | unclassified bacterium 1_1_47 | Broad |
| 337 | unclassified bacterium 1_7_47FAA | Broad |
| 338 | unclassified bacterium 4_1_31 | Broad |
| 339 | unclassified bacterium 4_1_37 FAA | Broad |
| 340 | unclassified bacterium 5_2_54FAA | Broad |
| 341 | unclassified bacterium 6_1_37FAA | Broad |
| 342 | unclassified bacterium 4_1_31 | Broad |
| 343 | unclassified_Desulfovibrionaceae bacterium 3_1_6 | Broad |
| 344 | unclassified_Desulfovibrionaceae bacterium 4_1_30 | Broad |
| 345 | unclassified_Enterobacteriaceae bacterium 9_2_54FAA | Broad |
| 346 | unclassified_Erysipelotrichaceae bacterium 2_2_44A | Broad |
| 347 | unclassified_Erysipelotrichaceae bacterium 6_1_45 | Broad |
| 348 | Ureaplasma urealyticum parvum | JCVI |
| 349 | Variovorax paradoxus2 | JCVI |
| 350 | Veillonella AF186071 | JCVI |
| 351 | Veillonella sp. 3_1_44 | Broad |
| 352 | Veillonella sp. 6_1_27 | Broad |
| 353 | Weissella paramesenteroides | BCM-HGSC |